Research Areas
Research Overview
Human fungal infections cause 2.5 million annual deaths yet remain neglected diseases. Our lab studies pathogens in the genus Cryptococcus, which are environmental fungi found in diverse ecological settings worldwide. Cryptococcus neoformans, the most prominent pathogen in this group, is an opportunistic yeast that causes life-threatening meningitis and is responsible for 19% of global HIV/AIDS deaths. Members of the Cryptococcus gattii species complex are related fungi of significant concern due to their ability to infect immunocompetent individuals. Our research aims to understand the unique biology of these fungi, with the long-term goal of developing new treatment and prevention strategies.
Genetic tools to study fungal pathogens
Advances in fungal genetics have laid the groundwork for rapid progress in dissecting fungal pathogenesis, yet there are numerous opportunities to expand this toolbox. A major goal is to develop and implement new methods that will open previously intractable biological questions.
Systematic functionalization of novel fungal genes
How do we study uncharacterized genes in divergent organisms? Cryptococcus species are members of the phylum Basidiomycota, which diverged from model fungi in the phylum Ascomycota (e.g., Saccharomyces cerevisiae) at least 450 million years ago. More than 1 in 5 C. neoformans genes lack readily identifiable orthologs outside of the basidiomycetes, making this species a valuable system to explore the diversity of eukaryotic gene function. Paradoxically, C. neoformans is also a powerful model for exploring conserved biology, as it retains cellular pathways that have been lost from traditional model yeasts yet are otherwise found broadly across the eukaryotic tree of life. Our lab uses high-throughput genetics and systems biology approaches to define cryptococcal gene function in an unbiased fashion, allowing us to study both unique pathogen adaptations and fundamental eukaryotic biology.

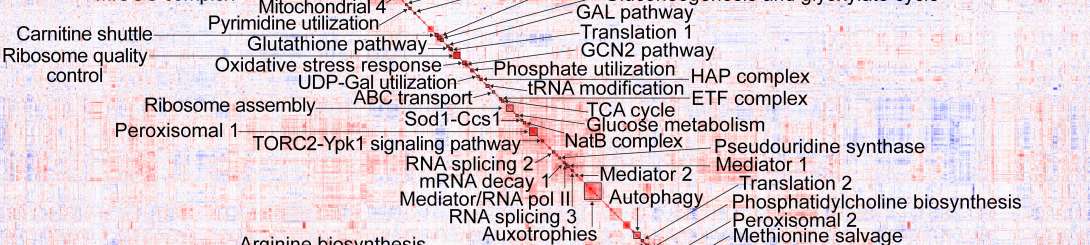
Figure caption: Mutant cofitness matrix clustering 1909 C. neoformans genes into functional modules. Boucher et al. (2025) Cell 188:4003-4024.
Molecular and evolutionary drivers of fungal pathogenesis
How do environmental microbes evolve to infect mammals? Many modern pathogens coevolved with their hosts for thousands of years, yet Cryptococcus species are environmental yeast that are not thought to have done so. We seek to understand how these fungi have retooled adaptations from the environmental lifestyles for mammalian infection, with an emphasis on dissecting the functions of novel virulence factors.
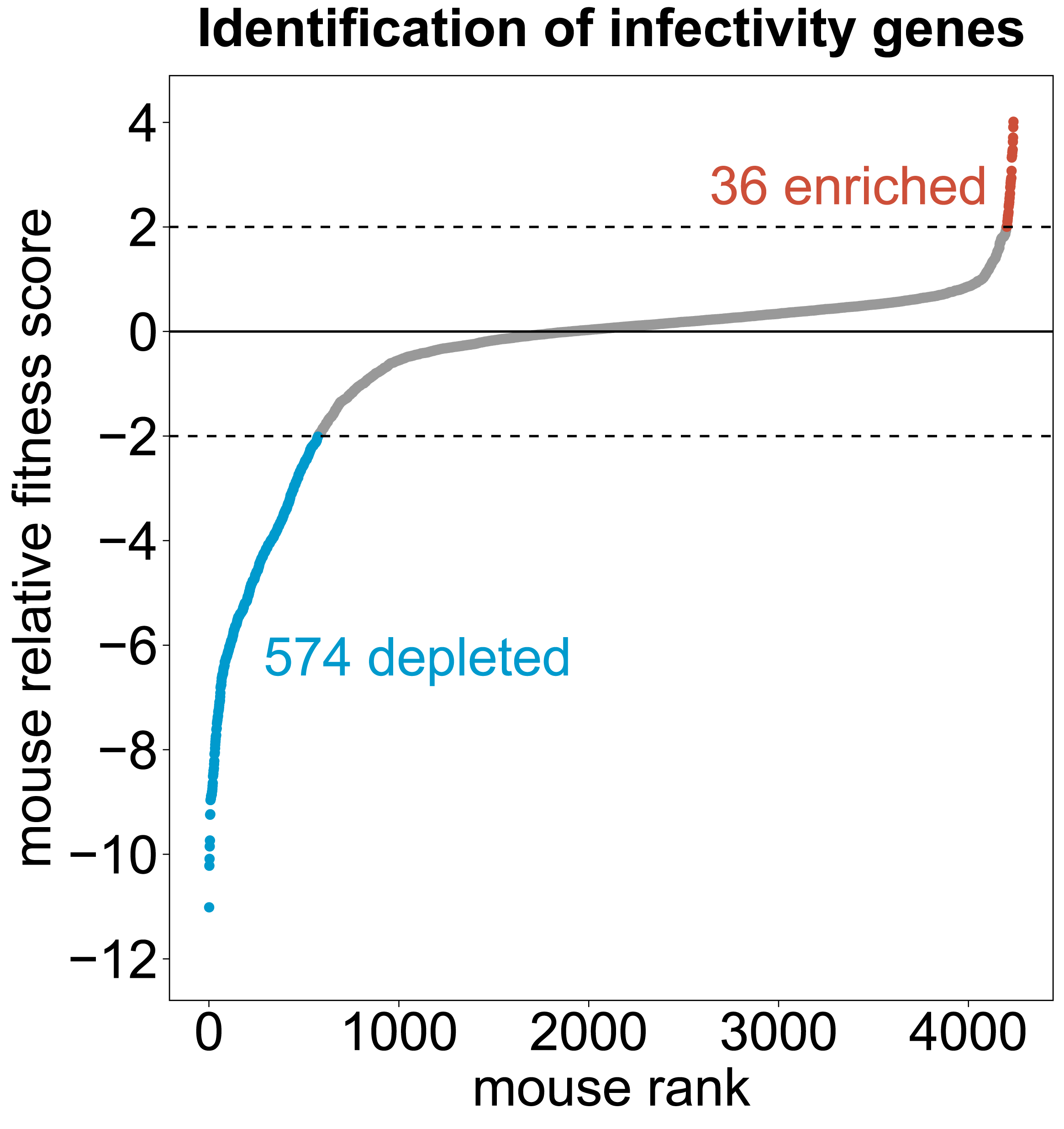
Figure caption: Identification of C. neoformans genes required for pulmonary colonization of C57BL/6J mice. Boucher et al. (2025) Cell 188:4003-4024.