Tuberculosis - The Tortoise
Would you be surprised to know that tuberculosis (TB) was the leading infectious disease killer in the world in 2019? Estimates of deaths attributed to TB were second only to COVID-19 in 2020 (roughly 1.5 million versus 1.8 million reported deaths, respectively). In the ongoing race to be the microbe responsible for the largest number of deaths across the planet, “hares” like HIV and SARS-CoV-2 explode onto the scene and appear to leave everyone else in the dust. But the number of deaths from HIV has declined over time with better prevention and treatment. And mortality from SARS-CoV-2 has also declined with widespread vaccine coverage and improved treatment options. As SARS-CoV-2 moves toward endemicity, the slow and steady “tortoise” in this race, Mycobacterium tuberculosis (MTB), may once again take over as the deadliest microbe on Earth.
One major factor to the persistence of MTB as a top ten infectious disease killer for thousands of years is the challenge of effective treatment. While the bacterium was only identified in 1882, the disease has been described in humans for more than 3,000 years and was even identified in human remains from 9,000 years ago. Furthermore, recognizing that MTB has a strong geographic component, especially as it relates to the ongoing spread of drug resistant strains, is key to understanding its worldwide impact on morbidity and mortality. Laboratory diagnostics and genomics are important newer components to determining effective treatment of TB cases.
Sequencing
Wadsworth Center scientists are on the forefront of identifying the fastest route to the best treatment regimen for each patient’s infection. In 2016, Wadsworth Center became the first state public health laboratory to perform whole genome sequencing (WGS) on clinical TB samples and six years later remains the only laboratory in the United States using WGS test results for clinical decision making. Since then, more than 4,000 TB genomes have been sequenced and hundreds of mutations associated with drug resistance have been detected. Using WGS to identify these genetic markers associated with drug resistance instead of traditional culture-based drug susceptibility testing (DST) improves timeliness – averaging seven days faster for the first line of drugs – and provides a highly comprehensive assessment of drug resistance. In an average year, 700-800 isolates are sequenced and roughly 10% have resistance to at least one drug. Being able to quickly provide these test results to clinicians so they can tailor the choice of medications for each case improves treatment for the individual and halts the spread of drug-resistant TB to others.
Collaboration
Considering the global impact of TB infection and the growing problem of drug resistance, it’s no surprise that researchers across the world are interested in learning more about how WGS can be used to accurately predict drug resistance. A connection between TB researchers at the University of Oxford and Wadsworth Center began with sharing of research findings and grew into joint research projects and an invitation to participate in CRyPTIC: Comprehensive Resistance Prediction for Tuberculosis: an International Consortium. This group of TB research institutions from all over the world is working “to achieve better, faster and more targeted treatment of multidrug-resistant TB via genetic resistance prediction.” Wadsworth Center contributed both WGS and DST data for 100 pan-susceptible TB isolates to the CRyPTIC project as well as WGS and DST data from additional isolates of interest. WGS data for another roughly 500 isolates have also been shared. The project now boasts the largest ever global dataset of clinical MTB isolates: over 15,000 samples from 27 countries on five continents.
Results
In addition to making the data available to the public, CRyPTIC is the primary author of nine publications describing advancements in the field and providing guidance for those who are implementing WGS for predicting drug resistance. Data were also shared with the World Health Organization to help create their first catalog of mutations known to be associated with drug resistance in MTB. At Wadsworth Center, scientists access the data to learn more about mutations previously identified in other areas of the world but newly found in New York. New collaborations with other members of CRyPTIC are ongoing and additional work to update the data could be down the road. In the meantime, Wadsworth Center scientists will continue to lead the way in the development and application of new TB diagnostics to benefit individual patients and the public’s health overall.
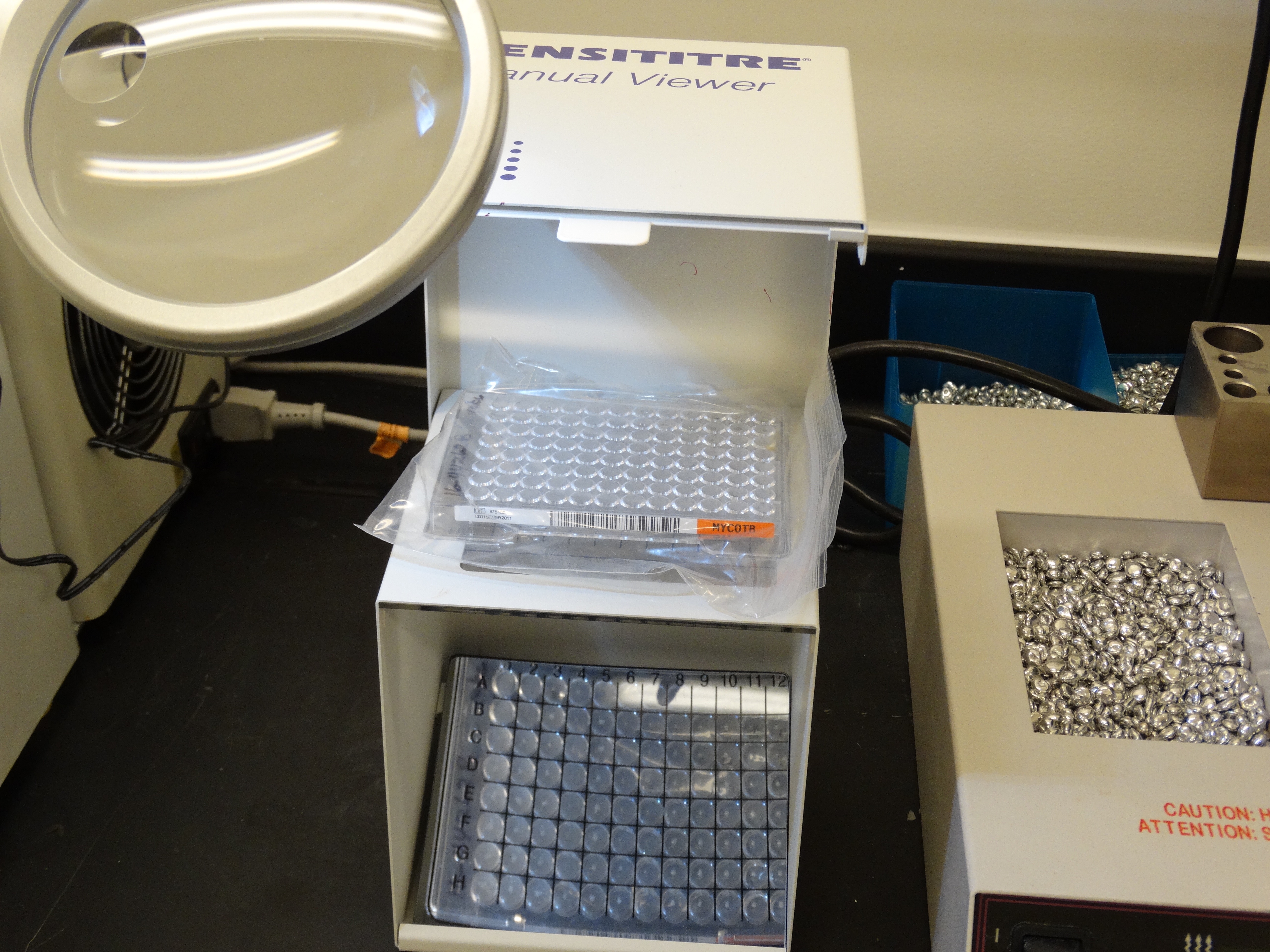
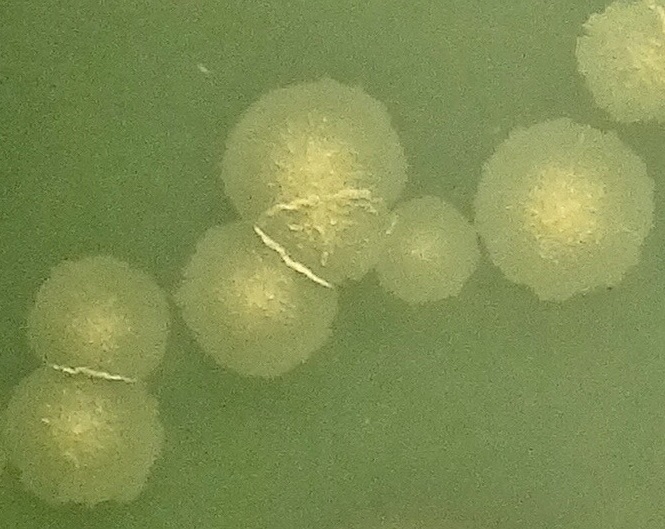

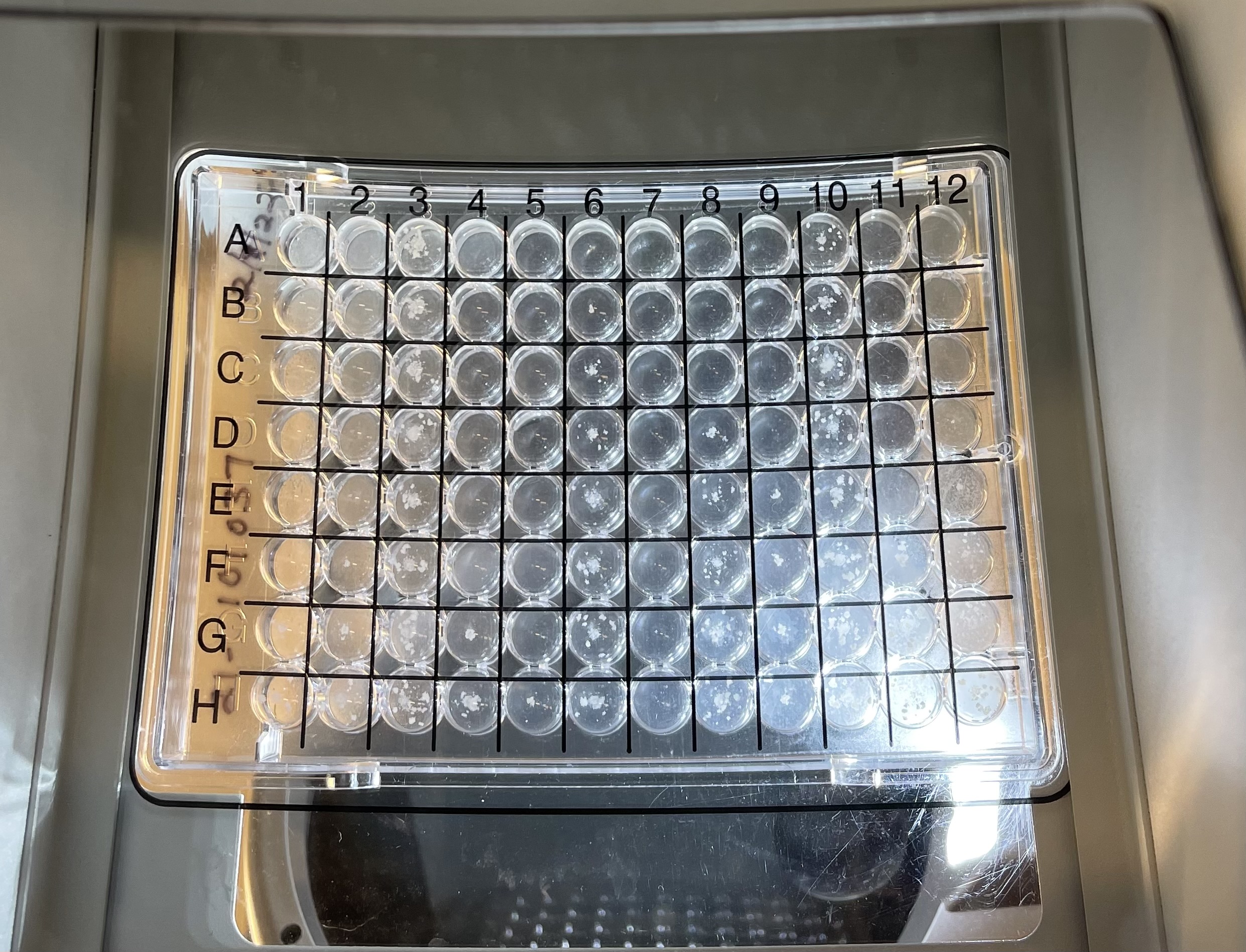
Left to right: Mycobacterium tuberculosis colonies on Middlebrook 7H10 agar, manual plate reader and 96-well broth microdilution plate.